based on lectures of Prof. Cimbala, ME341
The log-decrement method¶
The log-decrement is based on the following analysis:
and the damped natural frequency:
Using the output of the system in time (step function response) we need to solve for and simultaneously. The practical solution is the log-decrement method.
When , then the sine function is approximately and the magnitude only (peaks of the oscillating function) behave approximately as:
Therefore we plot the normalized step founction output minus 1, obtain a function that oscillates around zero, and try to extract the peaks. We can use only positive peaks and mark them as and their time instants, . From these values we can obtain:
The period of oscillations if we measure the time of cycles (e.g. in our example),
If we define the of the reduction of amplitude between each peak as :
, then the damping factor is recovered as:
and the rest is straightforward:
and
from IPython.core.display import Image
Image(filename='../img/log-decrement.png',width=600)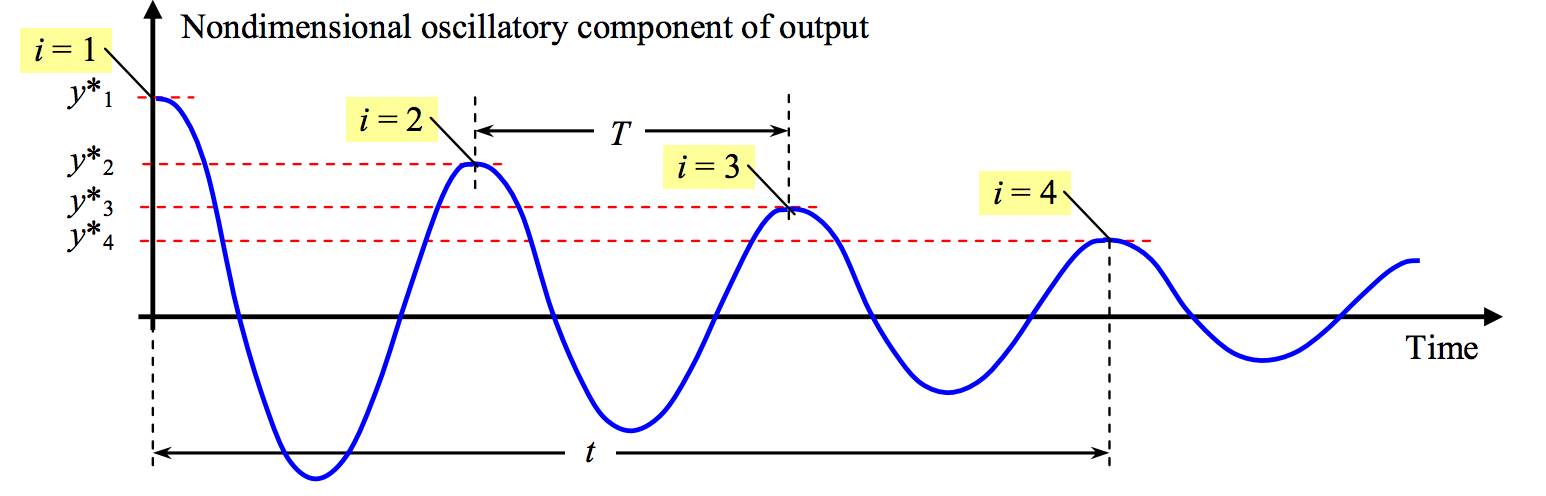
%pylab inline%pylab is deprecated, use %matplotlib inline and import the required libraries.
Populating the interactive namespace from numpy and matplotlib
/home/user/Documents/GitHub/mechanical-engineering-metrology-and-measurements/.venv/lib/python3.12/site-packages/IPython/core/magics/pylab.py:166: UserWarning: pylab import has clobbered these variables: ['sys']
`%matplotlib` prevents importing * from pylab and numpy
warn("pylab import has clobbered these variables: %s" % clobbered +
from scipy import signal
# Define transfer function
k = 1 # sensitivity
wn = 546.72 # rad/s
z=0.2 # damping
sys = signal.lti(k*wn**2,[1,2*z*wn, wn**2])
# step function output
t,y = sys.step(N=1000)
plot(t,y)
title('Step response')
xlabel('$t$ [sec]')
ylabel('E [V]')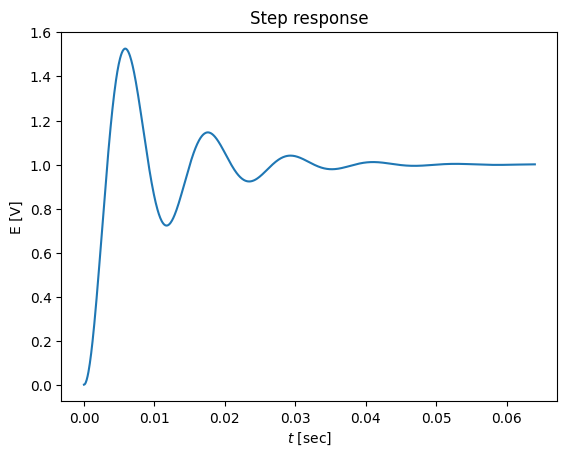
# note that sampling is sufficient, if not we need to apply the D/A reconstruction
# or interpolations, which will add more noise and uncertainty to the system identification# plot the data as a decrement
ts = t[::15]
ys = y[::15]
plot(ts,ys-1,'o')
title('Step response')
xlabel('$t$ [sec]')
ylabel('E [V]')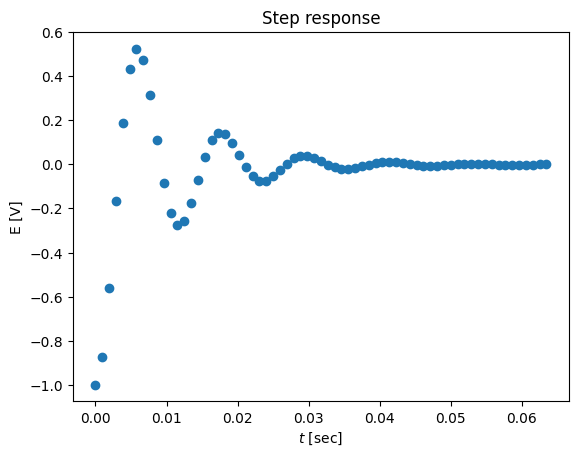
# we will use the open source peakdetect function from
def peakdet(v, delta, x = None):
"""
Converted from MATLAB script at http://billauer.co.il/peakdet.html
Returns two arrays
function [maxtab, mintab]=peakdet(v, delta, x)
%PEAKDET Detect peaks in a vector
% [MAXTAB, MINTAB] = PEAKDET(V, DELTA) finds the local
% maxima and minima ("peaks") in the vector V.
% MAXTAB and MINTAB consists of two columns. Column 1
% contains indices in V, and column 2 the found values.
%
% With [MAXTAB, MINTAB] = PEAKDET(V, DELTA, X) the indices
% in MAXTAB and MINTAB are replaced with the corresponding
% X-values.
%
% A point is considered a maximum peak if it has the maximal
% value, and was preceded (to the left) by a value lower by
% DELTA.
% Eli Billauer, 3.4.05 (Explicitly not copyrighted).
% This function is released to the public domain; Any use is allowed.
"""
maxtab = []
mintab = []
if x is None:
x = arange(len(v))
v = asarray(v)
if len(v) != len(x):
sys.exit('Input vectors v and x must have same length')
if not isscalar(delta):
sys.exit('Input argument delta must be a scalar')
if delta <= 0:
sys.exit('Input argument delta must be positive')
mn, mx = np.inf, -np.inf
mnpos, mxpos = np.nan, np.nan
lookformax = True
for i in arange(len(v)):
this = v[i]
if this > mx:
mx = this
mxpos = x[i]
if this < mn:
mn = this
mnpos = x[i]
if lookformax:
if this < mx-delta:
maxtab.append((mxpos, mx))
mn = this
mnpos = x[i]
lookformax = False
else:
if this > mn+delta:
mintab.append((mnpos, mn))
mx = this
mxpos = x[i]
lookformax = True
return array(maxtab), array(mintab)
# if __name__=="__main__":
# from matplotlib.pyplot import plot, scatter, show
# series = [0,0,0,2,0,0,0,-2,0,0,0,2,0,0,0,-2,0]
# maxtab, mintab = peakdet(series,.3)
# plot(series)
# scatter(array(maxtab)[:,0], array(maxtab)[:,1], color='blue')
# scatter(array(mintab)[:,0], array(mintab)[:,1], color='red')
# show()maxtab, mintab = peakdet(ys-1,.01,ts)# we need only positive peaks, maxima:
maxtabarray([[0.0057674 , 0.52586962],
[0.0173022 , 0.14414951],
[0.02979824, 0.03919356],
[0.04133304, 0.01110418]])# We see 4 peaks and therefore n = 4
tstar = maxtab[:,0]
ystar = maxtab[:,1]
# plot the data with the peaks
plot(ts,ys-1,'x',tstar,ystar,'ro',markersize=8)
title('Step response')
xlabel('$t$ [sec]')
ylabel('E [V]')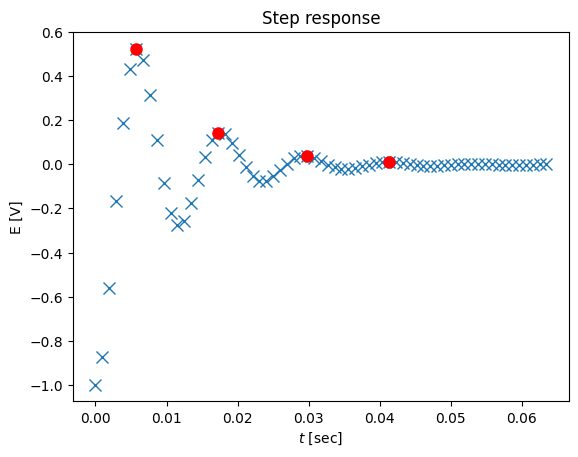
n = len(tstar)-1
print("cycles = %d" % n)cycles = 3
T = (tstar[-1] - tstar[0])/(n)
print ("period T= %4.3f sec" % T)period T= 0.012 sec
# delta
d = log(ystar[0]/ystar[-1])/(n)
print ("delta = %4.3f " % d)delta = 1.286
# recover the damping and the frequency:
zeta= d/(sqrt((2*pi)**2 + d**2))
omegad = 2*pi/T
omegan = omegad/(sqrt(1-zeta**2))
# output
print ("natural frequency = %4.3f" % omegan)
print ("damping factor = %4.3f" % zeta)
print ("compare to the original: 546.72, 0.2")natural frequency = 540.979
damping factor = 0.201
compare to the original: 546.72, 0.2